Automating experimentation with single-agent hypothesis selection and robotic evaluation
In this work we describe the implementation of a toy example for automated, iterative experimentation using Strateos and an autonomous hypothesis selection and evaluation agent written in python.

When it comes to experimentation the process is usually iterative with cycles of hypothesis generation, observation collection and hypothesis evaluation. One can see this prominently during the process of lead optimization in the medicinal chemistry field. Iterative experimentation seeks to explore a parameter space of a system such as a chemical structure and understand how this space correlates to an observation space such as the IC50 of that chemical structure interacting with a protein of interest.
Typically these input and output spaces can be extremely large so it is either technically impossible or financially infeasible to explore these spaces in a single experiment. In some cases an approach called Design of Experiments (DOE) can be used to perform statically driven experiments using schemes such as fractional-factorial designs to explore a space of factors the experimenter deems relevant. In lot's of cases these factors or features may not be known up front and naïve approaches for exploring these complex multidimensional spaces can be employed for a multi-parameter optimization in the case of gradient descent.
Looking more closely at lead optimization, typically a chemist will design a chemical structure that they believe will engage a target such as a protein, they are designing a 3D chemical topology to complimentarily engage with the 3D chemical topology of a protein. Of course there are lots of unknowns about a system like this, possibly too many to know up front, and how to translate that into a molecular design. Tools like virtual screening and docking can be used to help narrow down the hypothesis space into more valid hypotheses that should be evaluated but still the chemist must iterate through real world steps of chemical synthesis and assaying the material.
This approach can essentially be boiled down to experiments by rounds where a single round is as follows:
- Generate hypotheses about a system/phenomena
- Select a number of hypotheses to be evaluated
- Batch hypotheses together (hopefully) in single "experiment" or job
- Fetch all of the data and perform analyses in batch
- Evaluate each hypothesis to update mental model
- Begin the cycle again
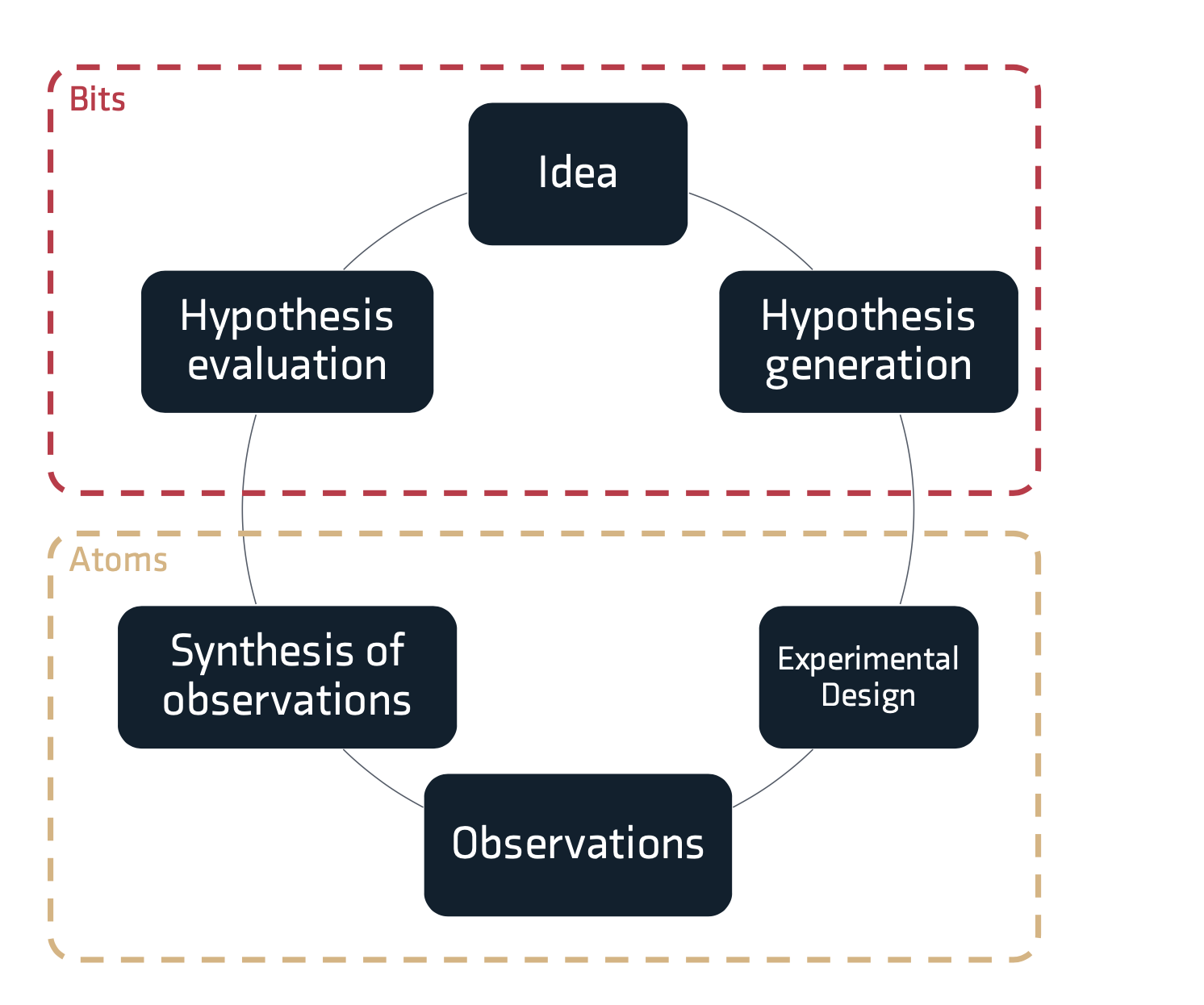
This structure is reflected in the overview of the scientific method in Figure 1 below. The top portion of the figure reflects the generation of ideas and hypotheses. In the world of bits or information, by using computational methods it is generally fairly easy to generate large volumes of hypotheses, however not all hypotheses are equally valid. For example, some hypotheses may have previously been evaluated, some hypotheses might not respect physical laws. If all hypotheses could be evaluated by experimentation with no scaling in level of effort, this would be a great position. Unfortunately hypothesis evaluation often requires physical experimentation using real atoms, with chemical and biological resources and the expenditure of a person's time. Given a fixed budget for hypothesis evaluation the experimenter must choose which hypotheses should be evaluated, this requires some rationale for picking sets of hypotheses.
In this work we describe the implementation of a toy example for automated, iterative experimentation using Strateos and an autonomous hypothesis selection and evaluation agent written in python. This approach can be extended to encompass a variety of hypotheses, experimental plans and analyses for a multitude of applications. Our goal for this simple example was to demonstrate the implementation of a system that automatically picks hypotheses for excitation and emission wavelengths for a specific fluorophore, generates experimental plans in Autoprotocol, creates a run via the Strateos API for execution, downloads the spectrophotometry data, and finally generates the next set of emission and excitation wavelengths to be tested. Through doing this, the agent will explore a space of excitation and emission wavelengths seeking the most efficient pairing.
Hypothesis generation and selection
An overview of the system architecture is shown in Figure 3. For component A of figure 3 we built a simple Python server that could generate and submit experiments to the Strateos web API and automatically fetch, analyze and submit the next experiment to Strateos. Though obvious, given prior knowledge, our macro-hypothesis of the system, is that there exists a most optimal excitation wavelength and emission wavelength pairing where the fluorophore exhibits the highest quantum yield.
To demonstrate a closed-loop, agent driven workflow, a common fluorophore (Alexa Fluor 594) was optimized for use with Strateos's spectrophotometry hardware. We optimized the fluorophore against Phosphate Buffered Saline (PBS) as a negative control and completed the entire process without human intervention. Each run was automatically submitted to the Strateos cloud lab platform and micro-well plates were stored in integrated, temperature controlled storage between runs.
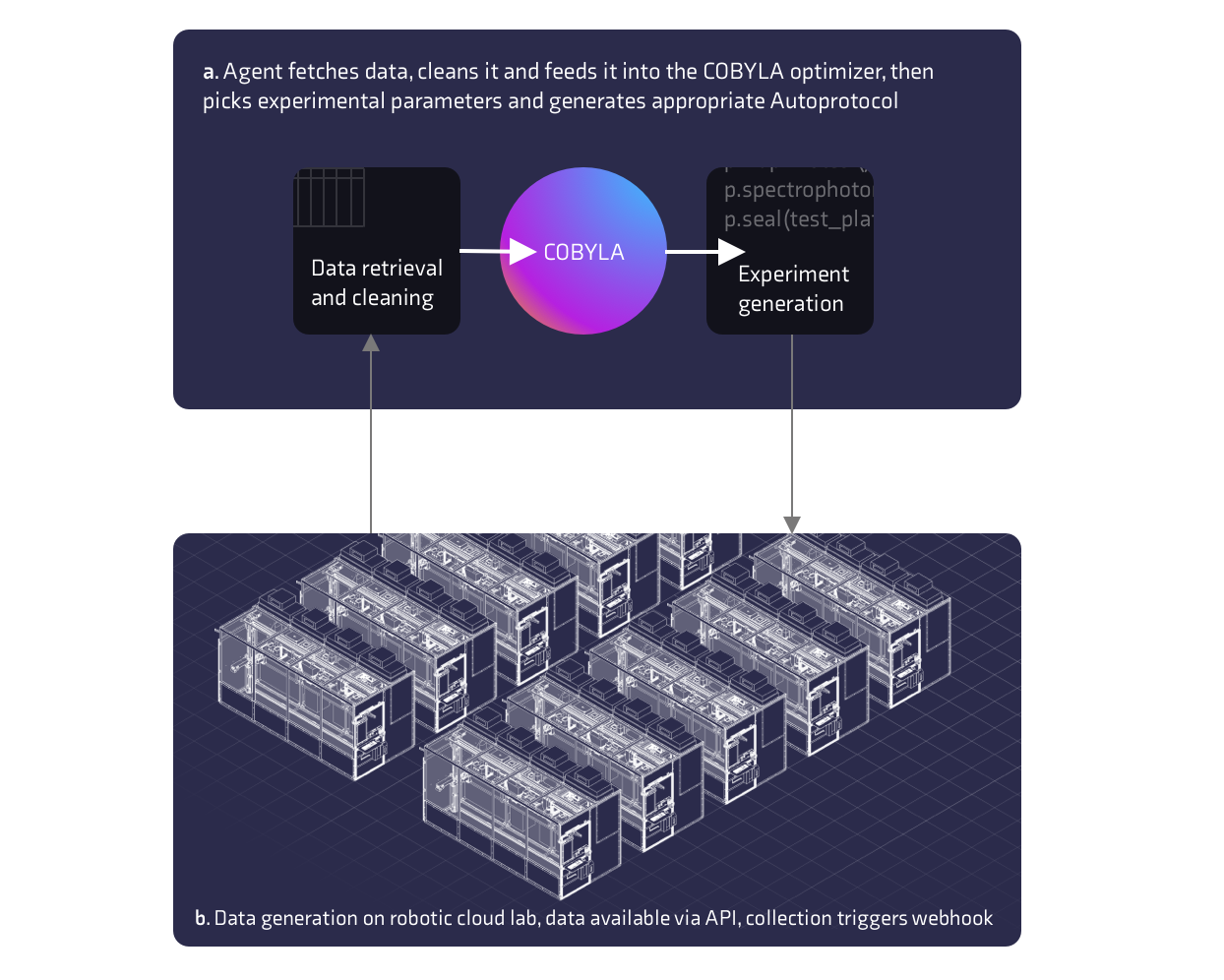
For the first round of experimentation the model randomly picked initial parameters to start. For subsequent rounds the parameters were sampled from the model. The problem was modeled as a constrained non-linear minimization problem, with an objective normalized ratio of positive and negative control read intensities. A constrained trust-region algorithm (COBYLA) was then applied with heuristics such as multiple local starts. Following the completion of experimental iterations, runs, the agent would receive a HTTP POST webhook event from the Strateos platform instructing the agent that the experiment had completed. The webhook infrastructure was set up simply with a single POST endpoint written in the open source framework Flask. Following notification, the agent would then fetch the spectrophotometry data from the Strateos web API, transform the data and use this to refine the linear model of the system for the next candidate solutions. After selecting the candidate solutions the agent would then generate a JSON object of Autoprotocol instructions for the next experiment and submit this to the Strateos Robotic Cloud Lab.
COBYLA
For the optimization piece of the agent we chose to implement a simple optimization algorithm available to everybody from the Scipy python package called COBYLA. COBYLA stands for Constraint Optimization By Linear Approximation. The SciPy implementation of COBYLA was chosen because it is broadly available and open source in addition to being sufficient for modeling this emission - excitation system.
From Wikipedia:
It works by iteratively approximating the actual constrained optimization problem with linear programming problems. During an iteration, an approximating linear programming problem is solved to obtain a candidate for the optimal solution. The candidate solution is evaluated using the original objective and constraint functions, yielding a new data point in the optimization space. This information is used to improve the approximating linear programming problem used for the next iteration of the algorithm. When the solution cannot be improved anymore, the step size is reduced, refining the search. When the step size becomes sufficiently small, the algorithm finishes.
Experimental fulfillment by robots
As with all other experiment jobs submitted to the Strateos platform, the job enters the queue and waits for all of the required infrastructure to be available. The Strateos Common Lab Environment then schedules and distributes all of the tasks to the platform to execute the experiment. Physical execution proceeds similarly to the below video:
Results
The best scoring search
The agent started 5 independent searches. The best performing search is shown in Figure 3, where we saw a relatively efficient convergence to a pairing of (540:nm, 625.6:nm). The decreasing step size of the COBYLA approach can be seen as the iterations proceed from iteration 1 to iteration 24. Since the agent's finding, differed from the manufacturer’s recommendation of (590:nm, 617:nm), it was thought that a line-scan should be performed with the resulting data shown in Figure 4.
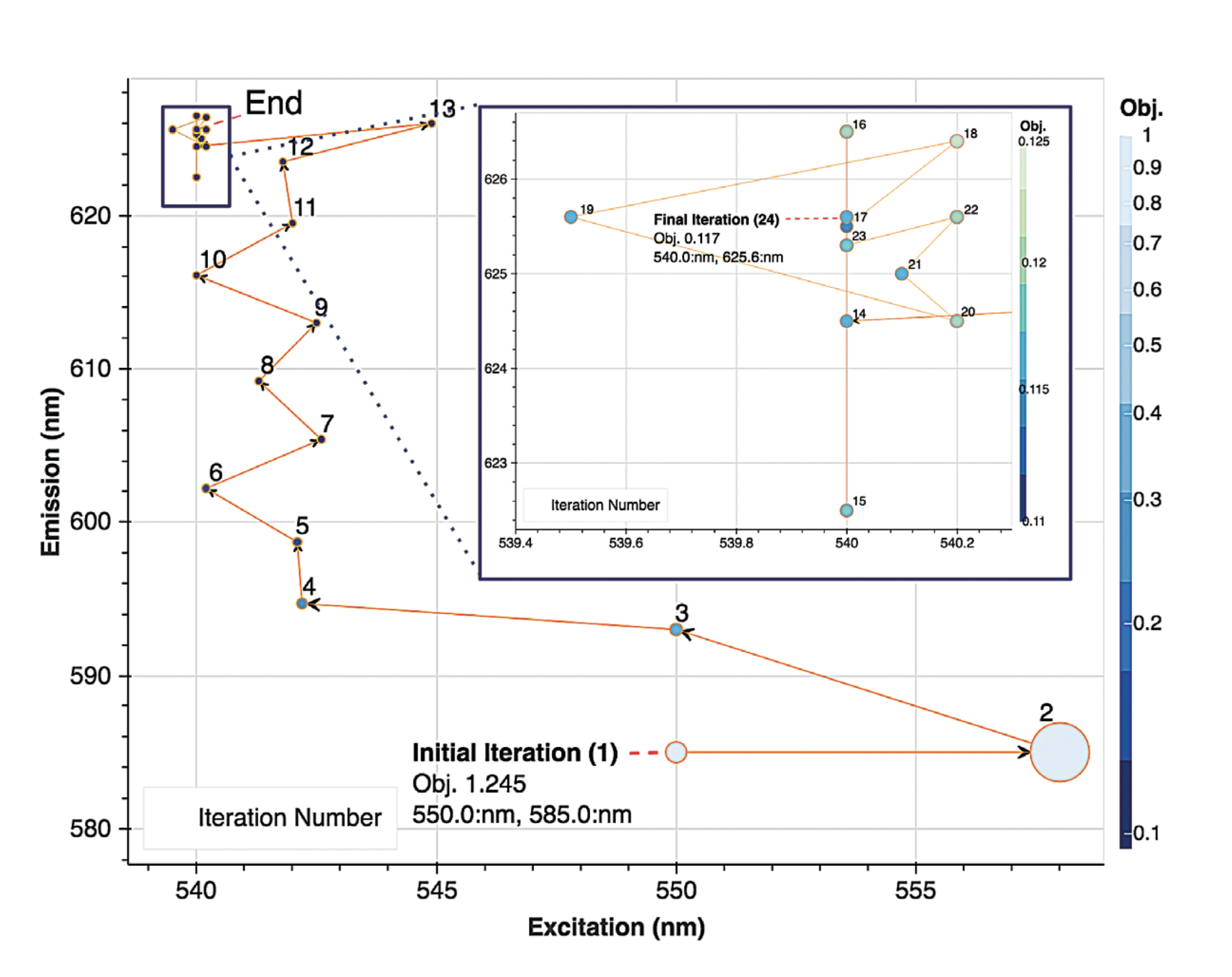
Figure 4 reaffirms our conclusion, where we saw that for the defined objective, the agent found a better minima, than the manufacturer guidelines.
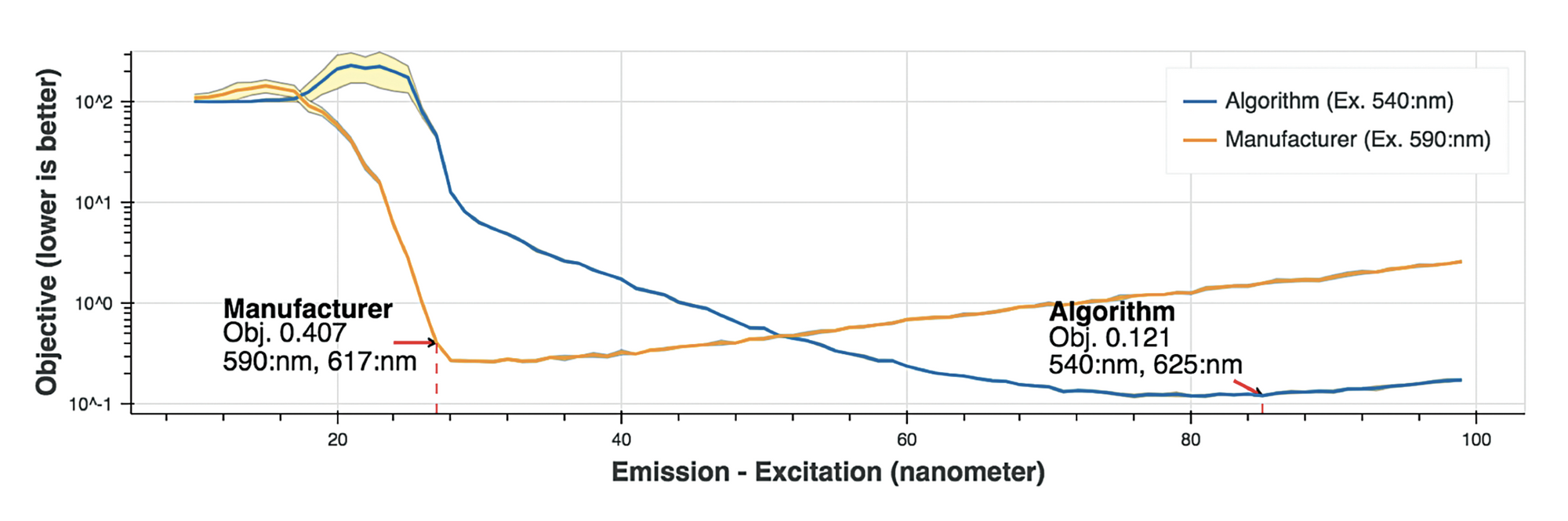
Verifying findings
To verify that the agent indeed found a global minimum, we conducted a coarse 5:nm scan of the landscape which is shown in Figure 5. This reaffirmed our results, showing that some of the traces did indeed get trapped in local minima, but our best trace (Figure 3) found the global minimum.
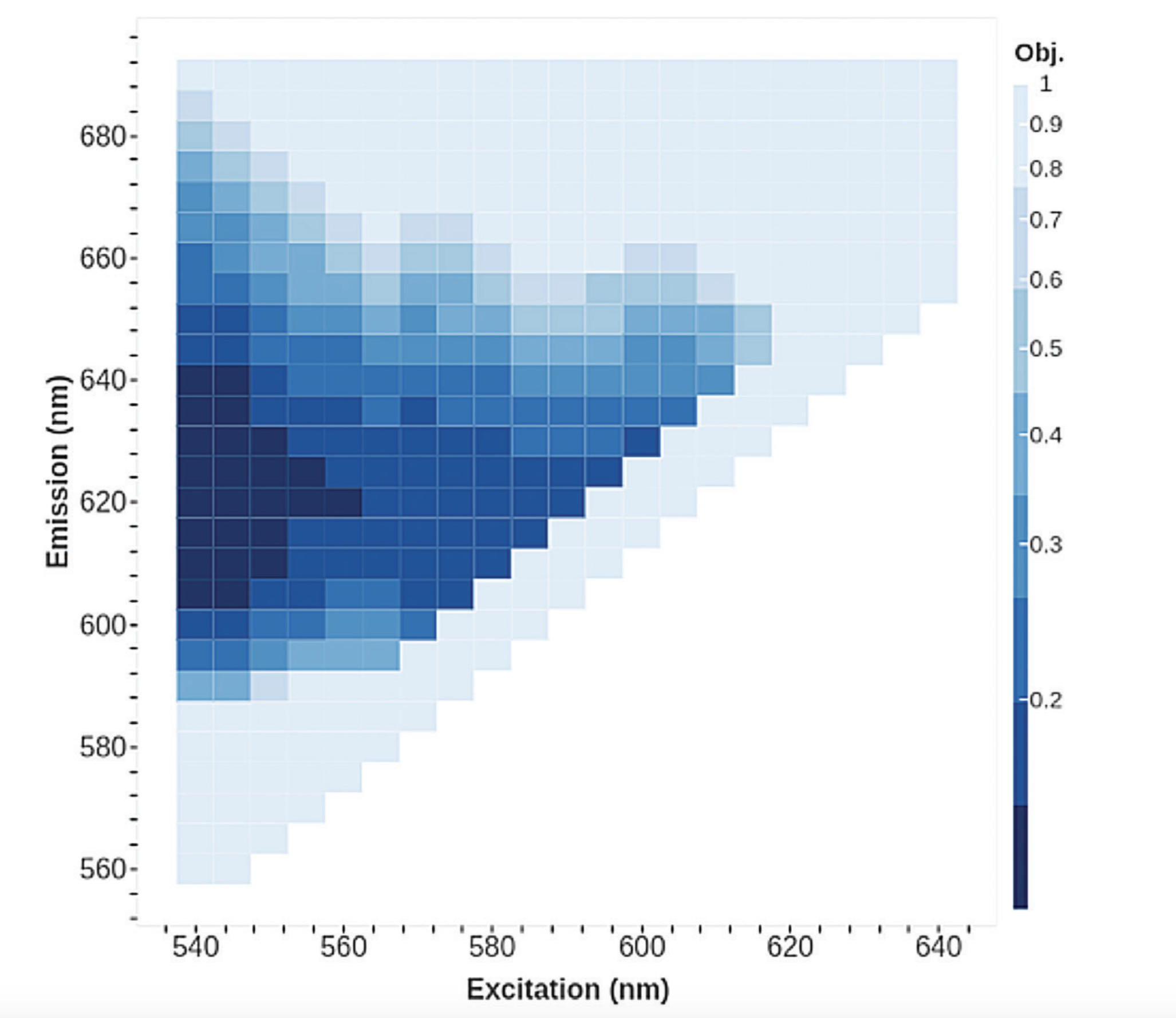
The agent using COBYLA with heuristics, took a total of 2.5 hours and 106 reads, as compared to a theoretical brute force search, which would have required 803103 reads and >3000 hours to search the entire space
Examining the overall time and reads taken, the agent using COBYLA with heuristics, took a total of 2.5 hours and 106 reads, as compared to a theoretical brute force search, which would have required 803,103 reads and >3000 hours to search the entire space. This translates to a 1000-fold improvement in search time, significantly decreasing the likelihood of reagent degradation effects. Overall, our approach lead to a better optimum than manufacturer recommendations likely due to device and reagent variations.
Looking forward
In this work we have shown how off-the-shelf, open source machine learning approaches can be plugged into the Strateos Robotic Cloud Lab Platform to create self exploring, scientific, automated agents. These agents can be applied to exploring complex spaces with too many factors for humans to explore alone, both physically and intellectually. We anticipate, and intend to demonstrate, that applying this approach to other aspects of system exploration and assay development will lead to a dramatic reduction in the time it takes to develop models of a system, enabling the acceleration of studying phenotypes, structure activity relationships and assay development.
If you're excited by working on automated scientific discovery, we're hiring.
Authors
Yang Choo, Donald Dalton, Ben Miles and Peter Lee.
